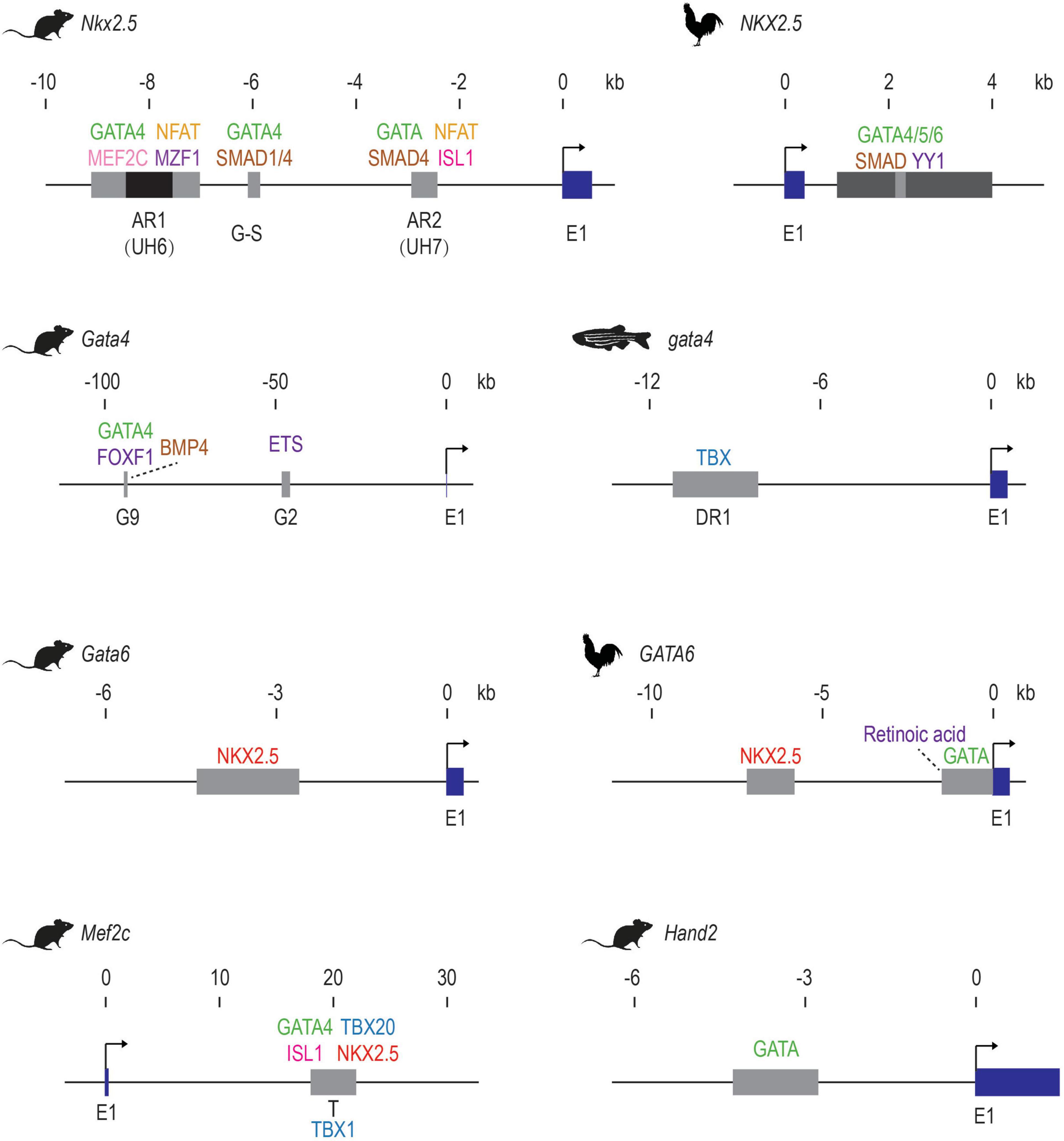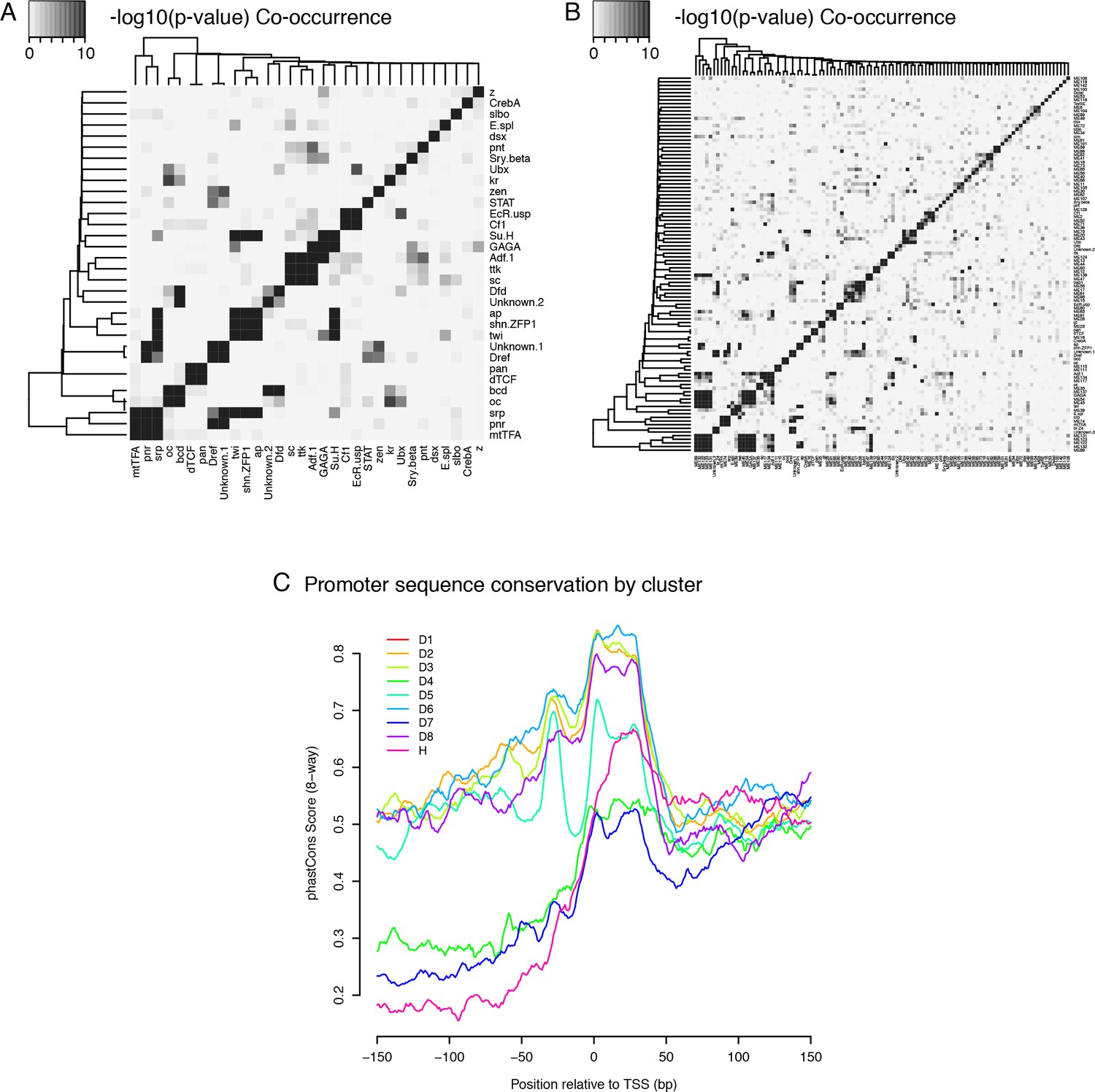
Conserved noncoding transcription and core promoter regulatory code in early Drosophila development | eLife
The miR-430 locus with extreme promoter density forms a transcription body during the minor wave of zygotic genome activation

The first murine zygotic transcription is promiscuous and uncoupled from splicing and 3′ processing | The EMBO Journal

The miR-430 locus with extreme promoter density is a transcription body organizer, which facilitates long range regulation in zygotic genome activation | bioRxiv

High-Titer Production of the Fungal Anhydrotetracycline, TAN-1612, in Engineered Yeasts | ACS Synthetic Biology
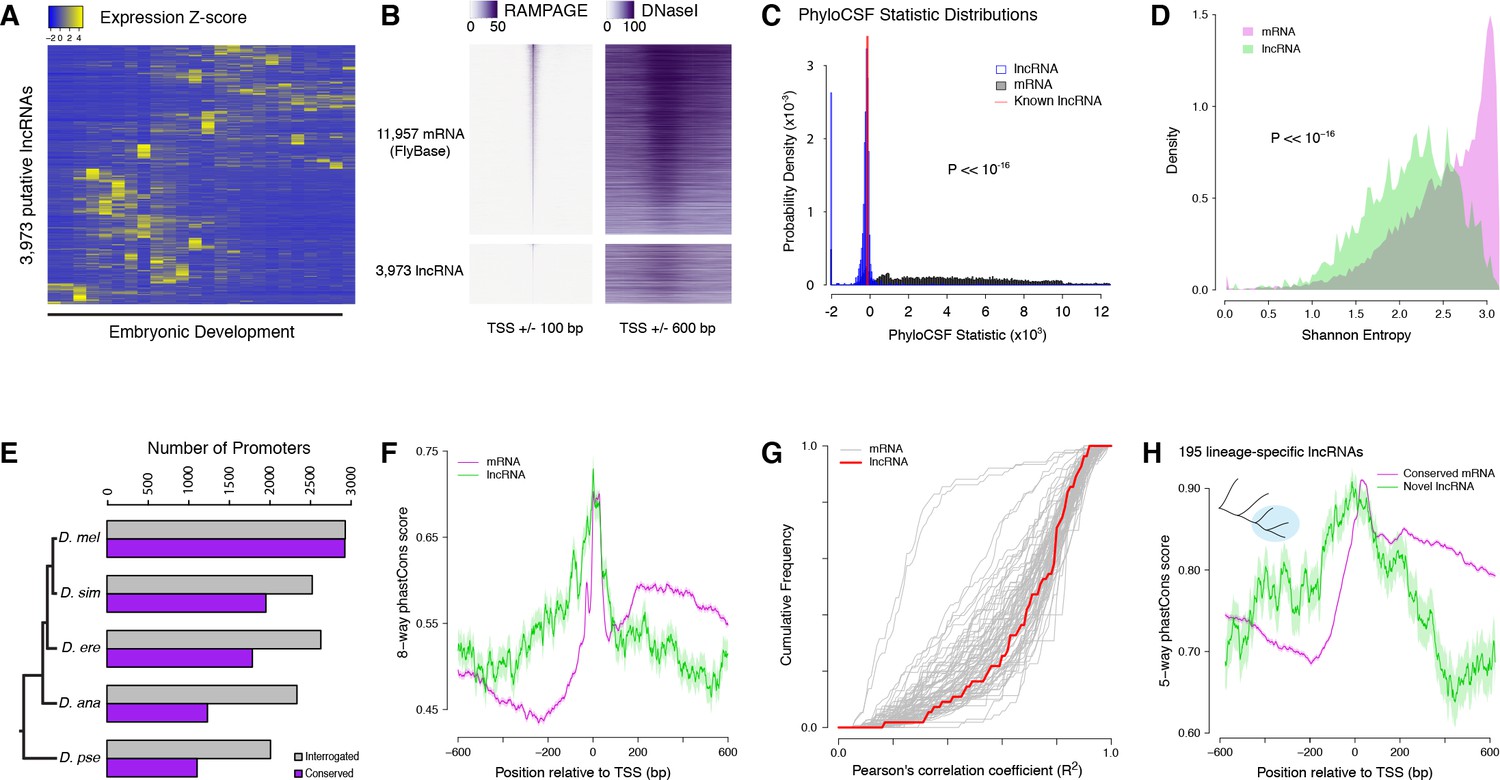
Conserved noncoding transcription and core promoter regulatory code in early Drosophila development | eLife

Conserved noncoding transcription and core promoter regulatory code in early Drosophila development | eLife

Gerhard KALINKA | Researcher | Dr. | Bundesanstalt für Materialforschung und -prüfung, Berlin | BAM | Department of Materials Engineering (Dpt.5) | Research profile

High-Titer Production of the Fungal Anhydrotetracycline, TAN-1612, in Engineered Yeasts | ACS Synthetic Biology
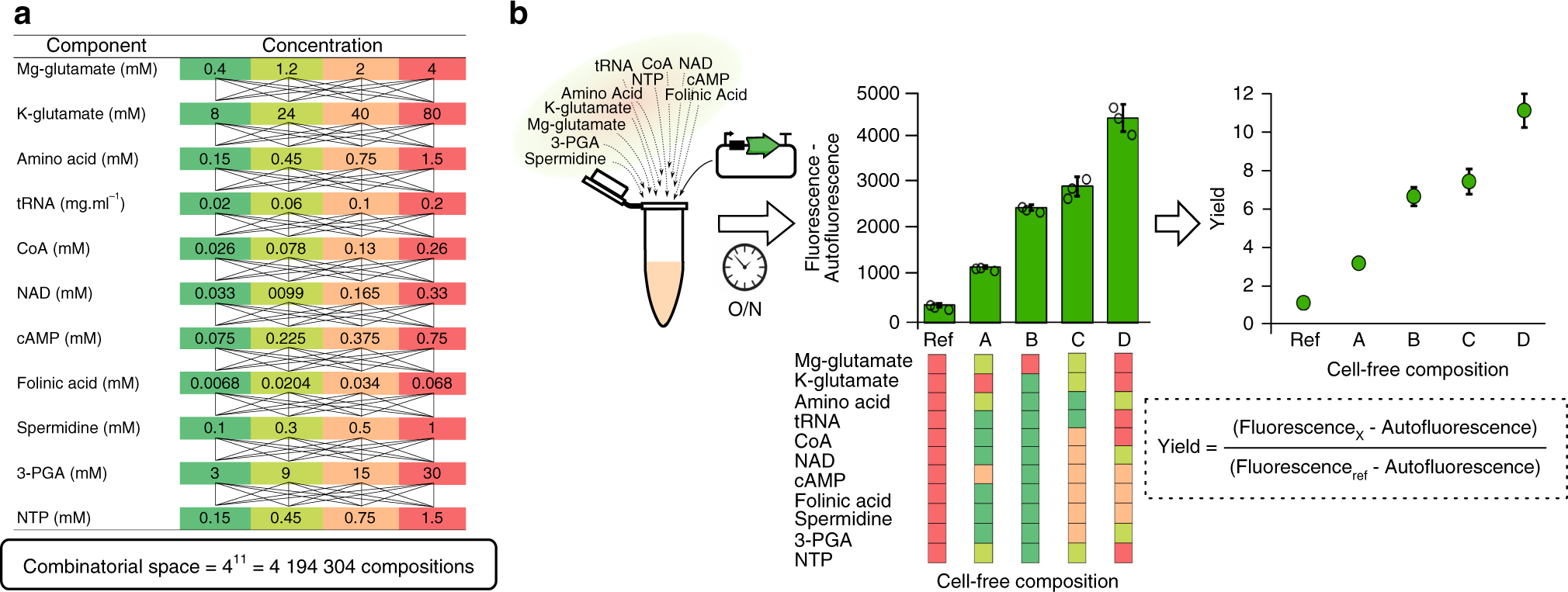
Large scale active-learning-guided exploration for in vitro protein production optimization | Nature Communications

In vivo measurements reveal a single 5′-intron is sufficient to increase protein expression level in Caenorhabditis elegans | Scientific Reports